-
EKHAM the Comics
What are “simulations” in advanced research? Is High Performance Computing the Holy Grail of scientific simulations? Let’s find out together through this unique Comic book
-
-
Keep up to date with the latest news from E-CAM
-
E-CAM Survey of Application Software
European Centre of Excellence
Supporting HPC simulations in industry and academia through software development, training and discussion in simulation and modeling
Quick Access Links
E-CAM Issue of Comics&Science
A Comic book about simulation, modelling and HPC.
Case Studies
Case studies and success stories related to pilot projects focused on industrially oriented problems
Software Repository
Access the software modules that have been documented by E-CAM
Attend an event
Our full calendar of events, that are part of the CECAM programme
Online Training Portal
Access the content captured at our ESDWs
Scientific Publications
Dissemination of E-CAM results in international peer-reviewed journals
E-CAM's stories
Interviews, opinion pieces, success stories, case studies.
Issue 16 – April 2021
E-CAM Newsletter of April …Challenges to Industry of drug substance development
Computation based methods …Development of an HTC-based, scalable committor analysis tool in OpenPathSampling opens avenues to investigate enzymatic mechanisms linked to Covid-19
The E-CAM HPC Centre …Proof of concept : recognition as a disruptive technology
Abstract The transformation …
View All Stories

Modules of the month
What is a module?

March
EESSI-based GitHub Action for Continuous Integration
Description This module sets up the European Environment for Scientific Software Installations (EESSI) for use in GitHub Workflows. The …March Module of the Month: DL_MESO (DPD) on Kokkos for enhanced performance portability
This work relates to the implementation of a performance portable version of DL_MESO (DPD) using the Kokkos library. It focuses …
View All Modules of the Month
Upcoming E-CAM events
Recent Posts

E-CAM Final Report
E-CAM started in October 2015 and received funding from …
Issue 16 – April 2021
E-CAM Newsletter of April 2021 Get the latest news …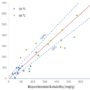
Challenges to Industry of drug substance development
Computation based methods play a growing role in all …
Development of an HTC-based, scalable committor analysis tool in OpenPathSampling opens avenues to investigate enzymatic mechanisms linked to Covid-19
The E-CAM HPC Centre of Excellence and a PRACE team …
